“Barely slept last night after we announced funding for a £3.2m DNA study partnership that I Chair. Seven years+ of advocacy and lobbying final paid off.” Sonya Chowdhury, CEO of Action for ME – a leader in the effort
When the Brits do it, sometimes they really do it. In 2005, they ladled out 5 million pounds for the largest and still the single most expensive study ever done in chronic fatigue syndrome (ME/CFS) – the notorious PACE trial. That, as we all know, didn’t turn out so well.
Fifteen years after the Brits funded the biggest ME/CFS clinical trial, they funded the largest and one of the most expensive ME/CFS studies ever: a $3.2 million pound ($4 million dollar) 20,000 person, 4-year genetic study.
As good as this study is, the willingness of the U.K. funders to plop down an large amount of money on a biological study for ME/CFS might be the most important takeaway.
Simon McGrath, in his blog “UK spends £3 million on the world’s biggest ME/CFS study“, pointed out that of the two funders of the study, the Medical Research Council (MRC) has only funded a few fairly small biomedical studies in ME/CFS before, and the National Institute of Health Research has never funded any.
Things can change – even in the U.K.
“As someone living with ME/CFS, I’m well aware that the patient community has waited a long time for a study such as this one that has such a strong, genuine element of patient involvement. All of us involved with this research project hope that it can start to address the totally unwarranted stigma and lack of understanding that so many patients with ME/CFS face on a daily basis.” Andy Devereux-Cooke
It also bears noticing that the London Times article on the study spent a significant amount of print on the relationship between COVID-19 and ME/CFS. The COVID-19 pandemic has the potential to forever change how people think about ME/CFS and there are signs that it already is. (Stay tuned for a blog on that coming up.)
The Study
The 20,000 person GWAS, or Genome-Wide Association Study, is a basic, ‘get down to the bones of our DNA’ study. The goal is to find single nucleotide polymorphisms (SNPs) – small changes in the genetic makeup of genes – which can change their functionality and contribute to ME/CFS.

“GWAS, or Genome-Wide Association Studies, are responsible for the deluge of discoveries in terms of the genetic risk factors for common disease that have been pouring out of research labs recently … if you’re successful…it allows you to zero in on a place in the genome that must be involved in disease risk without having to guess ahead of time what kind of gene you’re going to find.
The beauty of GWAS is it got us past the candidate gene approach, which had been pretty frustrating because most of the candidates turned out not to be right, to be able to say the whole set of genes are your candidates, let’s consider all of them, and here’s a strategy that’s comprehensive enough to allow you to do that.” Francis S. Collins, M.D., Ph.D.
That inclusivity has lead GWAS studies to become known for the sometimes weird, but deeply informative, results that have popped out.
The very first GWAS study done back in 2005 on macular degeneration identified 2 SNPs in genes encoding complement factors in the immune system which weren’t even remotely on anyone’s radar. In fact, the complement system was so off the charts in macular degeneration that complement research struggled to gain a foothold. Since then, though, two more complement genes have been identified, and drug development is under way.
A GWAS study uncovered the totally unexpected link between blood type and the risk for severe COVID-19 illness.
GWAS Studies: the Pluses and Minuses
“At every fork in the road … your genome has its thumb on the scales, making it slightly more likely that you will develop in one direction or another.” Eric Turkheimer, a behavioral geneticist
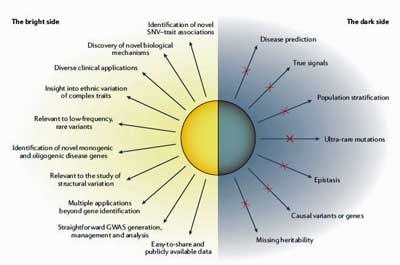
Fifteen years and about 4,000 GWAS studies later, we know a lot more about what GWAS studies can and can’t do. A recent review, “Benefits and limitations of genome-wide association studies” concluded that GWAS studies have “successfully identified risk loci for a vast number of diseases and traits”.
They’ve also often implicated “genes of unknown function or of previously unsuspected relevance” which after further study, have uncovered “novel biological mechanisms”. The data gained from GWAS studies can also form the basis for numerous other genetic studies.
While GWAS studies can provide incredible insights they tend to find gene variants that account for only a modest proportion of the heritability of complex traits. One reason for this is that it’s just very hard to identify some types of variants (common variants with very small effects, rare variants with small effects, ultra-rare variants) which, when they are all added up together, can significantly contribute to heritability.
Many traits in the body are polygenic; i.e. they arise from the interactions of many genes. While it would be nice, indeed, to highlight one or two or three genes that have turned the tables on people with ME/CFS, that’s not likely to happen. Instead of pointing to X gene causing a disease, GWAS studies tend to uncover arrays of genes and biological pathways that merit further study; i.e. they’re more likely to point fingers rather than directly pinpoint causes.
One geneticist reported that: “What GWASs do is plant a flag in the landscape and say “There’s something interesting here”. Next comes “the follow-up work to determine what that interesting thing is”.
The creators of DecodeME reported that they expected to find “dozens or hundreds of DNA differences that tilt the balance one way or another, changing someone’s risk of having ME/CFS.”
Once the DecodeME researchers find those genes, they’ll try to determine common pathways that have been disrupted in ME/CFS. DecodeME reported that they chose to study 20,000 people because other studies of this size generally found around five causal links between DNA and a disease.
The study’s 20,000 person size was chosen because other studies of that size have commonly uncovered 5 causal links to disease.
Our understanding of the role genetics plays in the development of ME/CFS at this point is pretty paltry – another reason for this study. A Utah family/genealogical study found “a strong support for a heritable contribution to predisposition to Chronic Fatigue Syndrome.” Several twin studies have suggested a genetic link to fatigue and/or ME/CFS exists as well.
Nothing even remotely like this has been attempted before in this disease. Given the rather meager stabs in the dark from the past genetic studies in ME/CFS, there’s no telling what this big exploratory study will find. A cluster of SNPs whacking the HPA axis or perhaps the energy metabolism, or the cardiovascular system or perhaps like the macular degeneration study, something totally unexpected.
Two mysterious SNPs found in Crohn’s disease provide an example of how potentially fulfilling the GWAS results can be. The two gene variants researcher found impaired the removal of damaged cellular components (autophagy). The plethora of damaged cellular goods, in turn, impaired the clearance of intracellular bacteria – and it was those intracellular bacteria which were causing the chronic inflammatory state in Crohn’s.
Prior to the GWAS study, no one dreamed autophagy might be a possible problem in Crohn’s.
Foundational Genetic Study – Every Disease Needs One
Getting at the potential genetic risk factors for a disease is the kind of foundational study that every disease needs.
DecodeME is scheduled to begin work in September and begin recruiting early next year. Its huge size means DecodeME may be bumping up against something Suzanne Vernon at the Bateman Horne Center warned against – the need to recruit enough real ME/CFS patients to fill the big studies that are going to start showing up in this disease.
The ME/CFS megastudy – presents a test for DecodeME – and the ME/CFS community – as DecodeME attempts to fill an ME/CFS study that’s magnitudes larger than any that have been attempted before.
The study will use a variety of methods to get across the goal line including PR campaign in early 2021 that includes press releases and TV and radio interviews; a patient-led social media campaign on Facebook, Instagram, and Twitter; digital advertising; and engagement with advocacy groups, and, of course, word of mouth.
Sonya Chowdhury gets the challenges that lie ahead:
“Simply put, we cannot do this without the determination and support of people with ME/CFS. Recruiting the 20,000 people we need is challenging – but absolutely achievable, by working in partnership with the CureME Biobank, charities, patient advocates, local support groups and others. People with ME/CFS can register their interest right now on the DecodeME website.”
So far, DecodeME is off to a good start with over 3,000 people registering as of yesterday.
Anyone with ME/CFS aged 16 years or over from the U.K. who wants to take part in the DecodeME study, or find out more, can register their interest now at http://www.decodeme.org.uk. Actual study recruitment will start next year, but let’s get the pool of possible participants filled to the brim to get this study off to a good start. DecodeME reported that
“Participants will be diagnosed using the CureME questionnaire that has been in use for the UK ME/CFS Biobank for several years. This is addition to participants reporting that they have been diagnosed with ME or CFS by a clinician. Participants simply need to answer a questionnaire and provide a saliva sample to be in the study.”
- Check out and sign up to get updates about the study on beautiful DecodeME website
- Check out FAQs here
Update – Several media articles have popped up on the study including two from The Times and this superb “long read” on ME/CFS.







Hi Cort,
Thank you so much for your wonderful blog. I’m in the UK and just registered my interest to participate in the study.
The UK Times have published relevant articles today and yesterday. Unfortunately with lazy journalism the articles inadvertently contribute to the historical narrative for ME/CFS, citing it as ‘controversial’.
That said the study is definitely a step in the right direction. Sending very best wishes and hope for good health to you and everyone following this fantastic blog. X
Thanks Rachel. A big congratulations to the researchers and the advocates involved. I don’t know how they managed – what arguments they used – to so totally flip the MRC and the other group – but it worked. 🙂
The fact that UK Prime Minister Boris Johnson seems to be developing ME/CFS post covid might explain such change… Several newspapers have talked about his need to take long naps every day, his intellectual sharpness is now gone, his quick jokes are no longer, his pale face, etc.
No way, Isa! Thanks for sharing that.
And thank you for the great article as always, Cort!
Cort… this has been worked on for well over a year. The study was mooted and a grant proposal was worked on many months before Covid-19, and the success in landing this big fish is down to the persistence and hard work of the CMRC under Stephen Holgate, and the steering and support of Jeremy Hunt. Absolutely nothing to do with Boris’ illness.
You imply that this success In convincing the MRC and the NIHR to finally fund biomedical research into ME/CFS may we be in some ways due to the controversy surrounding the “Pace trial and the energised advocacy community”. In my experience, it was the shrill and at times virulent elements in the community that delayed meaningful the engagement with government research fund allocators. But we all hope that these controversies and accusations can be relegated to the past, and that this funding marks the start of a new dawn with both government and private sources of funds!
Thanks Alex, I knew I was charting danger by stepping into the complicated world of ME advocacy in the UK.:)
Iwas careful to note that I really didn’t know how it happened, when I brought up the possibility that an energized advocacy community that emerged from the PACE trial controversy may have helped. I hadn’t considered that it could have hurt as well, which is retrospect, is entirely possible. I’m in full agreement that “advocates” who are more interesting in venting their anger than moving forward strategically have hurt us a great deal. You’re certainly closer to the action and know better than me – thanks for your input.
I didn’t mean to imply that this proposal was done in response to the COVID-19 problem and I apologize to anyone who got that impression. I well know that these things take a lot of dedicated work and time to put together. I saw the pre-COVID video, I think it was, which discussed the project, before writing the post.
I brought up the possibility that the funding environment may have improved given the link between problems with recovering from COVID-19 and ME/CFS. Again, I could be completely off base but given the huge and atypical funding this project received and the media attention the COVID-19/ME/CFS possible connections I think that’s a reasonable possibility to consider. One reason I brought it up is that things are beginning to happen for ME/CFS in the U.S. – blog coming up on that – which which wouldn’t have happen but for the COVID-19 / ME/CFS connection.
However, it happened the DecodeME project is great news, and I share your hope this is a start of a more fruitful, science-based collaboration between the UK government and ME/CFS researchers.
Alex,
I read your comment and two words stood out for me – ‘shrill’ – (piercing and high pitched) and ‘virulent’ (poisonous, malignant, bitter).
Given the circumstances and experiences associated with this illness, I think the way in which the situation is handled over the next while – including the publication of the new NICE guidelines in December 2020, could be instrumental in how the ME/CFS community reacts.
When you write: ‘But we all hope that these controversies and accusations can be relegated to the past…’ – I don’t know who the ‘we’ is but whoever you are, you may want to assess the language and approach you employ, if you don’t want to inflame an already incensed group of people.
Hi Cort, great to read your blog. I’m also in the UK. My 27 year old son with severe ME/CFS signed up for the study yesterday. I have a background in medical research and I follow this field closely. I must say I am very impressed by the way you combine detailed and accurate research information along with approachability in your writing.
Thanks Nicky! Good luck with your son.
Thank you so much. I really hope you are able to help those of us who suffer from this terrible disease xxx I am in the 25 percent severe range. Unfortunately I live in Australia so I am not able to partake in this study. I wish you all the best xx
Fingers Crossed this will lead to a cure xxx
Hopefully the insights the study opens up will be applicable to all of us. I would put ME/CFS patients in Australia, with their often British pedigree, high on the list of people likely to benefit the most from this study.
insertions
in the telomeres
are not limited
by nationality
talk intro is in Dutch and then switches to English:
https://m.youtube.com/watch?v=MCPjGJVuZz0&feature=youtu.be
we all bleed the same color…
and have same endings on telomeres….
pretty amazing findings jive
above link needs new edit 🙁
new link for Prusty lecture (in English after first minute)
https://m.youtube.com/watch?v=6jxMbrOwOPg#
“Why should we learn more on HHV-6 and HHV-7? A Perspective from ME/CFS. Dr. Bhupesh Prusty”
on virus insertion into telomere;
impact on mitochondria
Hello Cort. To me, having been diagnosed with this dreadful disease a few years ago, I am both excited and hopeful that some very positive results will come out of this study. I am in my low 70’s , born in Britain but now living in New Zealand. Perhaps New Zealand with its history of British immigrants is another area which will hopefully reap also the benefit of your study. My personal thanks go to everyone involved in trying to decode ME/CFS. All the very best of luck.
“We’re focusing on recruiting from the UK for now and are already well on our way with early sign-ups. But we’ll expand to other countries if necessary, so if you’re outside the UK, please do sign up now. You must be 16 or over to take part.”
Quote is from the DecodeME website (link is at the end of Cort’s article).
I’m in Australia too and I’ve just signed up – it’s a very simple process and you can de-register any time you like. “We’ll expand to other countries if necessary” implies this will happen if they don’t get enough people from the UK or maybe other reasons too. In any case, signing up at this point is like an expression of interest, so they know how many people are interested and so we can be kept updated on the project.
I’m in the uk, I signed up yesterday too although as I’m not a typical ME patient I dont know if I’ll pass the criteria. As a consultant who was supposed to be a specialist said ME maybe they’ll include me. They wont let recovered people participate so for the first time I’m glad that I’ve never managed to get back to normal. I dont know how big the ME societies mailing lists are but I do know that when I attended a conference people with ME were actually desperate to participate in any medical studies.
A lot of people are sceptical about handing over DNA to anyone – I am too but feel I have to do this.
I think this has been prompted by Covid-19 but maybe the psychiatric lobby are hoping it’ll find nothing.
Do they use the Oxford criteria? I read they look not at ME-patiënts but people with fatique. So i don’t expect anything of this study.
I was wondering about this. What would make sense would be to test post-infectious severe and very severe patients. No one else.
Gijs, on their website, they state ‘Those who pass the Canadian Consensus or Institute of Medicine criteria can participate.’
No, They definitely do not use the Oxford criteria. They use CCC And the Institute of medicine criteria. So please reset your expectations!
Oke, that is great news but they will never find 20.000 ME patiënts according to the CCC criteria. It would even surprise me if they find 1000.
This disease is much less common if they use the ccc criteria. It is all about fatique then you find 20.000 patiënts.
I signed up yesterday. I was impressed by the number of articles all over Facebook yesterday. I hope the promotion continues.
i signed up but I am mostly recovered. I still think they should use pwme who have mostly recovered in case they relapse and then they can take another DNA and see what slipped.
Finding “the gene” will not do a bit of good. Phenomenal waste of money, and time.
Some years back, they found the gene for progeria. Improvements since then? Zero.
They found the gene for cystic fibrosis . . . there are a hundred or more mutations. Progress since then?
They keep finding the gene for autism . . . one, two, and many, many genes. Progress following each of those discoveries? Zero.
And so on . . . many genes have been identified. But so what? Other than a very small number of success stories, so what?
Better they would investigate the chemistry more.
I think more than finding “the gene” which they have said is quite unlikely, is the that the study will identify pathways for more study. I think it will provide a piece of puzzle and we will need a lot more. So far, all this big data studies haven’t really fulfilled their promise – they point at general not specific areas. Let’s hope that changes over time
I am not super optimistic either. What I am more concerned about is the selection of patients. Years ago (15-20?) someone insisted on the need to identify sub-groups of patients. This was the way massive progress were made in the treatment of leukemia. Yet we still see all patients as a group, no distinction between trigger events, duration, severity, age at onset, those who sleep a lot, those who cannot, those with fibromyalgia, those without, those with EDS, those without, … The list is long. 10-20,000 patients seem a lot, but once sub-groups are identified one may end up with groups of just a couple of hundreds. Will that be enough? Perhaps researchers should “re-use” the same patients who took part in previous studies (research on brain inflammation in Japan a couple of years ago, immune profile in Stanford, etc.) as we already have detailed info on these people.
That’s fantastic that the UK is actually going to do this study! For those in the US, there is ‘All of Us’ where the NIH is collecting medical information (including genetic) on 1 million patients and will follow them over time.
Although not specifically for ME/CFS, the data can be used by any qualified researcher (the site is info.joinallofus.org) and almost any US citizen can join.
My concern with genetic studies is how practical interventions can be developed from wayward SNPS–unless you are talking about CRISPR. Maybe they can find issues in various metabolic pathways…
Unfortunately I hear (about my EDS), ‘It’s genetic, so there is nothing to be done.’ That’s wrong of course, but I think it may be a long time before this study bears fruit. Bad genes might even become the excuse for our illness.
However, it now appears that about 20% of Covid survivors have become ‘long-haulers’ in that they are experiencing their symptoms wax and wane for much longer than expected. As you have mentioned before Cort, that might be the instigation to pour money into post viral fatigue (of any sort)–and that might be a very good thing for us!
I don’t know how this study came about but I think the COVID-19 connection is going to result in a new era for ME/CFS. There’s just too much going in the media for post-viral illnesses to be ignored any more.
Nancy, glad you mentioned the US’s “All of Us” study currently reporting 350,000 participants to date of which close to 2500 have ME/CFS.
Anyone wishing to volunteer can do so at http://www.joinallofus.org. Data about the ME/CFS participants can be viewed here: https://researchallof us.org.
Francis Collins recently announced the “All of Us” study will be expanded to test the blood of 10,000 of its participants for COVID-19 antibodies (indicating past exposure to the virus) as well as to study in depth over time participants who contract COVID-19.
Hopefully this will catch COVID-19 “long haulers” who could then be compared to the ME/CFS participants already in the “All of Us” database.
Sorry for the typo– the website for the “All of Us” research data is: https://researchallofus.org
From what I understand, this study, if successful, may create a sort of map – with areas highlighted for further investigation. Seems like a really good place to start.
I agree with Anonymous that this study could lead nowhere really and be a complete waste of time. What they need to do is study the genes of severe ME if they want to do this study. Then everyone knows they are studying ME and not any CF or CFS. As far as I know, genes only tell you who is likely to get a disease. DNA is a double stranded helix and everyone has slight differences or faults I guess.
https://www.researchgate.net/publication/319157873_Contesting_the_psychiatric_framing_of_MECFS
I came across this article ‘Contesting the Psychiatric Framing of ME/CFS’ on Researchgate and thought it was a really comprehensive discussion of a very thorny subject.
From their Abstract they state ‘We highlight the potentially harmful consequences of psychiatrisation which can lead to people’s experiential knowledge being discredited.’
I know it’s slightly off topic but any mention of the UK and ME/CFS inevitably triggers the association with the PACE trial, as Cort mentions in his blog.
Anyway for me, it pretty much sums up many thoughts that circle in my head and it’s quite a relief to see those thoughts explored in print.